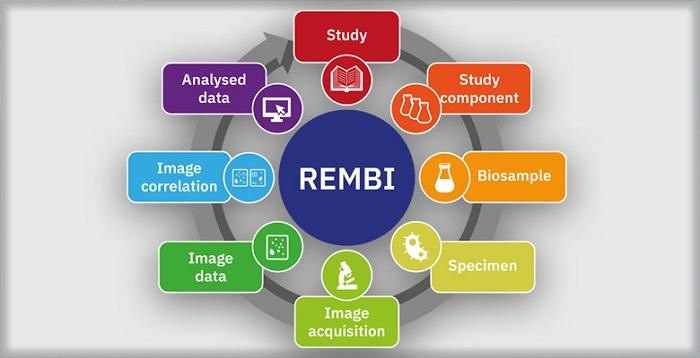
EMBL-EBI and collaborators propose a set of Recommended Metadata for Biological Images to help improve reusability
As the volume and quality of bioimages increases – and the technologies become more accessible through projects like EMBL’s Imaging Centre or Euro-Bioimaging – the scientific community also needs to be able to openly share the data and maximise reuse. This is important for applications like validating biological observations and developing machine learning methods. To do this, data producers require robust data infrastructure and standards, including metadata – details on why and how the images were produced.
A comment published in Nature Methods proposes a series of metadata guidelines called Recommended Metadata for Biological Images (REMBI). The guidelines have been compiled by specialists from EMBL’s European Bioinformatics Institute (EMBL-EBI) and colleagues at over 30 other institutions.
The guidelines are a result of a series of meetings with representatives from the different bioimaging communities, including light, electron and X-ray microscopy communities. The aim of the proposed guidelines is to provide a framework for discussing imaging data sharing. The hope is to reach community-wide consensus on the level of detail that is optimal.
The current version of REMBI is available online.
The standard will be adopted by the BioImage Archive, EMPIAR, Cell-IDR and Tissue-IDR databases. The authors hope that other archives will also adopt REMBI and engage with them to shape the future development of the standard in the spirit of FAIR data (Findable, Accessible, Interoperable and Reproducible).
Interested parties can contact the group on rembi@ebi.ac.uk.
Source article
SARKANS, U., et al. (2021). REMBI: Recommended Metadata for Biological Images—enabling reuse of microscopy data in biology. Nature Methods. Published online 21 05; DOI: 10.1038/s41592-021-01166-8